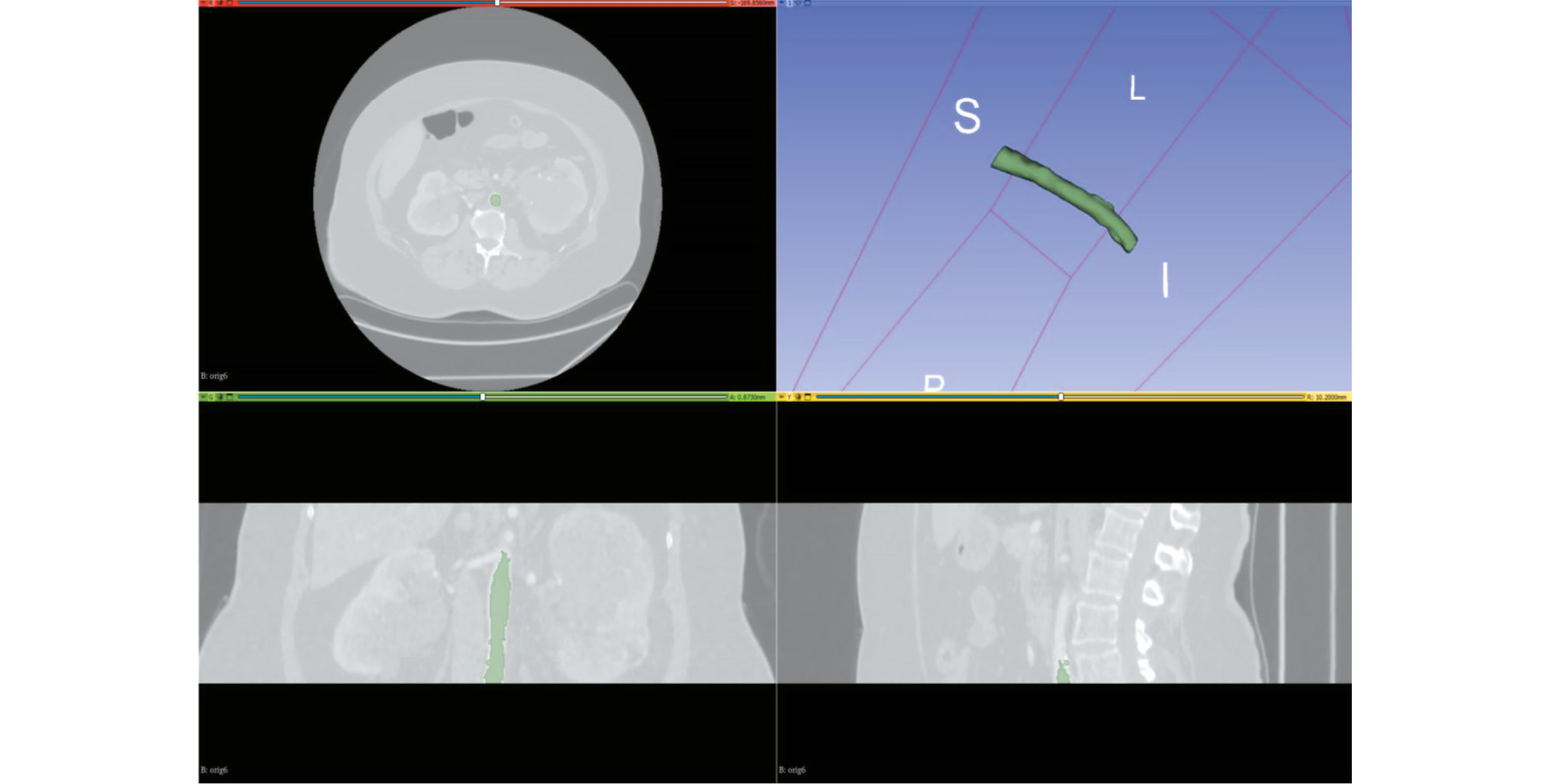
CT angiography dataset with abdominal aorta segmentation
- Authors: Kodenko M.R.1,2, Vasilev Y.A.1, Solovev A.V.1,3, Gatin D.V.1, Yasakova E.P.1, Guseva A.V.1,2, Reshetnikov R.V.1
-
Affiliations:
- Research and Practical Clinical Center for Diagnostics and Telemedicine Technologies
- Bauman Moscow State Technical University
- Morozov Children's City Clinical Hospital
- Issue: Vol 6, No 1 (2025)
- Pages: 23-32
- Section: Datasets
- Submitted: 02.09.2024
- Accepted: 29.10.2024
- Published: 25.03.2025
- URL: https://jdigitaldiagnostics.com/DD/article/view/635589
- DOI: https://doi.org/10.17816/DD635589
- ID: 635589
Cite item
Full Text
Abstract
BACKGROUND: Artificial intelligence algorithms are used to analyze images obtained through radiological diagnostic methods. The effectiveness of such algorithms depends on the availability of relevant and representative training datasets. The volume of such data in the public domain should be increased, particularly datasets containing abdominal aorta computed tomography angiography images, with pathology classification and vessel segmentation. The limitations of existing solutions include small sample sizes, restricted dataset specialization, and inconsistent dataset preparation methodologies.
Aim: To create an open dataset containing computed tomography angiography images of abdominal aorta segmentation for normal aorta, aneurysm, thrombosis, and calcification.
MATERIALS AND METHODS: A technical specification for dataset preparation was developed according to the methodology for testing artificial intelligence algorithms, the required sample size was calculated, and approval was obtained from an independent ethics committee. Regarding dataset creation, a previously developed original semiautomatic segmentation algorithm using Slicer 3D software was employed. The inclusion criteria were computed tomography angiography or abdominal computed tomography scans with contrast, arterial phase, and slice thickness ≤3 mm. Conversely, the exclusion criteria were presence of foreign bodies in the aorta lumen and aortic dissection. The algorithm was tested on patient data obtained from the Unified Radiological Information System. An expert evaluation was conducted to assess the compliance of obtained results with the established requirements and evaluate the time efficiency of using the developed segmentation algorithm.
RESULTS: The calculated sample size was 100 angiographic studies, including arterial phase scans with a slice thickness of ≤1.2 mm. Population data: number of unique patients, 100; percentage of female patients, 51%; and median age, 62 years (age range: 18–84 years). Pathology (including combined pathology) was detected in 61% of cases: 60 studies showed signs of calcification, 18 revealed aortic dilation, and 18 determined signs of thrombosed lumen. The average time to process one study (100 slices) using the developed segmentation algorithm was 0.8 hours.
CONCLUSIONS: A dataset containing 100 computed tomography angiography results with abdominal aorta segmentation for normal cases, aneurysm, thrombosis, and calcification was created. The dataset is publicly available and can be used for developing and testing artificial intelligence algorithms and for anthropomorphic modeling of the abdominal aorta.
Full Text
ОБОСНОВАНИЕ
Брюшная аорта представляет собой протяжённый сосуд, обладающий пациент-специфичной геометрией, структура и просвет которого могут видоизменяться при развитии различных патологий [1]. «Золотым стандартом» диагностики патологий брюшного отдела аорты является компьютерная томографическая ангиография (КТА) [2]. Её результаты используют не только для постановки точного диагноза, но и для планирования хирургического вмешательства [3]. Применение технологий искусственного интеллекта для обработки данных КТА позволяет автоматизировать ряд рутинных медицинских задач и, следовательно, оптимизировать процессы диагностики и лечения пациентов [4]. Обучение и тестирование алгоритмов искусственного интеллекта требуют наличия репрезентативных наборов данных [5]. Недостаток качественно размеченных результатов исследований лучевой диагностики в открытом доступе существенно замедляет развитие его клинически применимых алгоритмов. В рамках данной работы основное внимание уделено набору данных, содержащему результаты КТА с сегментацией брюшного отдела аорты для случаев наличия и отсутствия рентгенологических признаков её атеросклеротического поражения — расширение просвета, тромбоз и кальциноз стенки [6].
Обзор готовых решений в открытом доступе выявил малое число разнородных наборов данных, позволяющих лишь частично решить задачи обучения и тестирования алгоритмов искусственного интеллекта. Существующие решения недостаточно детализировано представляют описание методики получения и разметки изображений, полученных с помощью лучевых методов диагностики1, 2 [7–10]. Более того, часть исследований не содержат необходимого клинического и демографического описания пациентов, чьи результаты включены в набор данных1 [7]. Также стоит отметить одноцентровой характер собранных данных2 и узкую специализацию наборов данных [8], ограничивающую их использование в рамках конкретной клинической задачи.
ЦЕЛЬ
Создание открытого набора данных, содержащего результаты КТА с сегментацией брюшного отдела аорты для случаев нормы, расширения, тромбоза и кальциноза.
МАТЕРИАЛЫ И МЕТОДЫ
Дизайн исследования
Исследование проведено в соответствии с методологией MI-CLAIM [11], регламентирующей порядок клинического использования технологий искусственного интеллекта, а также с учётом правил проведения тестирования его алгоритмов в лучевой диагностике [12]. Дизайн исследования — ретроспективный анализ результатов КТА.
Критерии соответствия
Для включения в исследование установлены следующие критерии:
- результаты КТА органов брюшной полости или компьютерной томографии (КТ) органов брюшной полости с контрастированием;
- результаты КТА брюшной аорты и её ветвей;
- наличие артериальной фазы сканирования в проведённом исследовании;
- толщина срезов ≤3 мм.
Критерии исключения:
- признаки расслоения сосудистой стенки;
- наличие внутрисосудистых стентов (протезов).
Условия проведения
Результаты КТА получены без ограничений по медицинским организациям из единой радиологической информационной системы (ЕРИС), которая входит в состав государственной информационной системы в сфере здравоохранения субъекта Российской Федерации — «Единая медицинская информационно-аналитическая система города Москвы» (ЕМИАС)3.
Период сбора данных: с 24.03.2017 по 22.04.2022. Врач-рентгенолог с опытом работы более трёх лет проводил отбор результатов исследований вручную.
Методы обработки данных
Исходные данные представлены в формате DICOM (Digital Imaging and Communications in Medicine) [13], а результаты сегментации сохранены в формате NifTI (Neuroimaging Informatics Technology Initiative) [14]. Сегментацию изображений осуществляли с использованием программного обеспечения Slicer 3D (версия 5.0.2)4 по оригинальной методике, подробно изложенной в работах, которые мы ранее опубликовали [15, 16]. Сценарий разметки включал в себя следующие этапы:
- применение инструмента оконтуривания для области интереса на каждом пятом срезе (либо каждом третьем для областей выраженной извитости сосуда);
- последующее применение инструмента межсрезовой аппроксимации маски «fill between slices».
При необходимости проводили ручную экспертную корректировку полученной маски с помощью базовых инструментов «кисть» и «ластик».
Сегментацию изображений осуществлял врачрентгенолог с 3-летним опытом работы. В дальнейшем её результаты смотрел, верифицировал и, при необходимости, корректировал врач-эксперт с 10-летним опытом работы. Структура выходных данных имеет вид парных файлов «исследование и маска». Использованы различные маски сегментации для различного типа тканей: аорта, тромб и кальцинаты. Время разметки каждого результата исследования фиксировали для определения среднего времени на обработку одного изображения КТА.
Этическая экспертиза
Для работы с данными пациентов получено разрешение независимого этического комитета на базе ГБУЗ НПКЦ ДиТ ДЗМ (протокол №5/2023 от 25.05.2023). Выгруженные из ЕРИС результаты исследований деперсонализированы по стандарту DICOM: удалены персональные данные пациентов, а уникальный идентификатор исследования искажён.
Статистический анализ
Принципы расчёта размера выборки: целевое назначение набора данных — тестирование и обучение алгоритмов искусственного интеллекта. Для расчёта размера выборки использован метод оценки минимально необходимого размера при анализе метрик диагностической точности алгоритмов [17]. Параметры для расчёта: оценочная метрика площади под кривой (area under receiver operating curve, AUROC) с учётом допустимого доверительного интервала. Его ширина выбрана равной 10% [18], целевое значение AUROC установлено с учётом доверительного интервала так, чтобы нижнее граничное значение превышало порог, установленный методическими указаниями по тестированию алгоритмов искусственного интеллекта [12] в рамках «Московского эксперимента по внедрению компьютерного зрения» [19]. Для расчётов использован автоматический калькулятор presize5. Расчёт требуемого объёма выборки по выбранной методике показал, что минимально число исследований при балансе классов «норма» : «патология» 1:1 составляет ~80 результатов исследования, однако с учётом вероятной вариативности класса «патология» принято решение увеличить размер выборки до 100.
Методы статистического анализа данных: статистический анализ данных в рамках работы ограничен описательной статистикой включённых результатов исследований пациентов и структуры набора данных. Расчёт проведён с использованием средств MS Excel6.
РЕЗУЛЬТАТЫ
Объекты исследования
Популяционные данные:
- общее число пациентов — 100;
- доля пациентов женского пола — 51%;
- медиана возраста — 62 года при размахе значений от 18 до 84 лет.
Основные результаты исследования
Итоговый набор данных содержал 100 размеченных результатов КТА с общим объёмом 3,12 ГБ. Детализация структуры набора данных приведена в табл. 1. Баланс классов «норма» : «патология» в выборке из 80 результатов исследований составил ~1:1, дополнительно 20 результатов исследований содержали паттерны патологических изменений. Примеры разметки данных КТА, сегментации просвета сосудов и различного типа изменений (просвет, кальцинаты, тромб) представлены на рис. 1, 2. Структура набора данных показана на рис. 3. Набор данных зарегистрирован как база данных: свидетельство о государственной регистрации № 2024621990 [20] и доступен для свободного скачивания под лицензией Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported (CC BY-NCND 3.0)7.
Таблица 1. Структура набора данных
Патологические изменения | № исследования | ||||||||||||||||||||||||
1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 | 25 | |
Расширения брюшной аорты | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
Максимальный диаметр брюшной аорты, мм | 18 | 16 | 16 | 17 | 18 | 18 | 18 | 18 | 17 | 19 | 21 | 16 | 17 | 23 | 18 | 16 | 17 | 15 | 17 | 16 | 21 | 17 | 17 | 17 | 16 |
Кальцинаты | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 1 | 1 | 1 | 0 | 1 |
Тромбоз | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 1 |
26 | 27 | 28 | 29 | 30 | 31 | 32 | 33 | 34 | 35 | 36 | 37 | 38 | 39 | 40 | 41 | 42 | 43 | 44 | 45 | 46 | 47 | 48 | 49 | 50 | |
Расширения брюшной аорты | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
Максимальный диаметр брюшной аорты, мм | 16 | 23 | 20 | 20 | 15 | 21 | 15 | 18 | 18 | 29 | 22 | 11 | 15 | 37 | 18 | 19 | 17 | 16 | 21 | 15 | 15 | 24 | 18 | 17 | 18 |
Кальцинаты | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | 0 | 1 |
Тромбоз | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
51 | 52 | 53 | 54 | 55 | 56 | 57 | 58 | 59 | 60 | 61 | 62 | 63 | 64 | 65 | 66 | 67 | 68 | 69 | 70 | 71 | 72 | 73 | 74 | 75 | |
Расширения брюшной аорты | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 0 | 0 | 1 |
Максимальный диаметр брюшной аорты, мм | 19 | 19 | 14 | 34 | 40 | 35 | 37 | 20 | 37 | 19 | 36 | 35 | 19 | 33 | 37 | 32 | 43 | 23 | 30 | 27 | 23 | 27 | 19 | 16 | 41 |
Кальцинаты | 0 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
Тромбоз | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 |
76 | 77 | 78 | 79 | 80 | 81 | 82 | 83 | 84 | 85 | 86 | 87 | 88 | 89 | 90 | 91 | 92 | 93 | 94 | 95 | 96 | 97 | 98 | 99 | 100 | |
Расширения брюшной аорты | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
Максимальный диаметр брюшной аорты, мм | 33 | 18 | 20 | 19 | 18 | 14 | 16 | 17 | 22 | 20 | 14 | 14 | 17 | 18 | 21 | 13 | 17 | 18 | 15 | 16 | 12 | 16 | 16 | 14 | 23 |
Кальцинаты | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 1 | 0 | 0 | 1 |
Тромбоз | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
Примечание. 0 — наличие признака; 1 — отсутствие признака. | |||||||||||||||||||||||||
Рис. 1. Сегментация брюшного отдела аорты с помощью программного обеспечения 3D-Slicer.
Рис. 2. Разметка сосуда с помощью программного обеспечения 3D-Slicer: красный цвет — просвет сосуда; фиолетовый цвет — тромбы; жёлтый цвет — кальцинаты.
Рис. 3. Фрагмент набора данных, иллюстрирующий его организацию: файл classification.xlsx содержит перечень исследований, включённых в набор данных и результаты классификации. В директории studies находятся исследования и бинарные маски сегментации в формате NifTI — Neuroimaging Informatics Technology Initiative (.nii).
Дополнительные результаты исследования
К дополнительным результатам исследования можно отнести среднее время, затраченное на разметку одного среза с использованием предложенной методологии, которое варьировало от 25 с до 2 минут в сложных случаях, или усреднённо 0,8 часа на обработку результата КТА объёмом 100 срезов. Время сегментации представлено для оценки трудозатрат при повторении данного исследования или его масштабировании. Сегментация объектов позволяет использовать результаты работы для трёхмерного моделирования сосудов, востребованного, как показано ранее, при создании медицинских обучающих симуляторов и решении исследовательских задач в области гемодинамики [21, 22].
ОБСУЖДЕНИЕ
Резюме основного результата исследования
С использованием разработанной ранее методики создан набор данных, содержащий 100 исследований КТА с сегментацией брюшного отдела аорты для случаев нормы, расширения просвета, тромбоза и кальциноза стенки.
Обсуждение основного результата исследования
Поиск готовых решений по известным ресурсам набора данных открытого доступа, таких как Google datasets8, Kaggle Datasets9, Roboflow Universe10 и Images.cv11, демонстрирует небольшое число разнородных результатов. Так, L. Radl и соавт. [7] представляют набор данных, содержащий 56 результатов КТА аорты (1 случай аневризмы, 5 случаев диссекции и 50 случаев нормы) с разметкой дуги аорты и её ветвей, а также брюшной аорты с подвздошными артериями, при этом авторы сообщают, что исходные данные заимствованы из различных источников. Разметку провели с использованием программного обеспечения Slicer 3D и описали её методику, однако число разметчиков и уровень их квалификации не указали. Набор данных, предложенный L. Yaneth1, заявлен как результат нейросетевой сегментации брюшной аорты по результатам КТ. Он содержит мало сопроводительной информации — известно только то, что его подготовил и отсегментировал автор самостоятельно с использованием машинного обучения, в него включено 5289 изображений с разделением на 13 файлов с результатами исследований и масками. Данные о балансе классов набора данных, порядок разметки изображений и требования к разметчикам, так же как и популяционные данные, отсутствуют. В открытом доступе существует набор данных new-workspace-xlgzg2, содержащий 936 изображений в формате JPEG, на которых размечен контур брюшной аорты. Они представляют собой срезы различных изображений КТ без указания их параметров и информации о разметке. Дополнительный поиск по литературным источникам, ссылающимся на наборы данных в открытом доступе, с помощью ресурса Google Scholar по запросу «Abdominal aorta, computed tomographic angiography, segmentation, dataset», позволил дополнить результаты. M. Imran и соавт. [8] сообщают о разметке 59 результатов КТА брюшной аорты с разрешением 512×512 пикселей, толщиной среза от 0,8 до 2 мм и средним числом срезов в исследовании 734: отобраны данные пациентов с расслоением аорты из базы данных авторов. Сегментация аорты и её ветвей осуществлена аспирантами полуавтоматически с использованием программного обеспечения Slicer 3D, сведения о медицинской специализации и опыте разметчиков не представлены. A. Fantanzinni и соавт. [9] в своей работе сообщают о подготовленном ими наборе данных, содержащих сегментацию аорты и её ветвей по результатам 80 предоперационных КТА пациентов из университетского госпиталя San Martino (Италия), у которых диагностирована аневризма брюшной аорты (иные состояния аорты в набор данных не включены). Отмечено участие рентгенологов–экспертов при сегментации аорты с использованием программного обеспечения ITK Snap12. Набор данных не включает изображений с признаками нормальной аорты и иных патологий, кроме аневризмы, а работа не содержит прямой ссылки для его загрузки. Следует отметить, что сходное описание имеет работа Y. Jung и соавт. [10]. Авторы приводят подробное описание (без прямой ссылки) собственного набора данных, содержащего 60 результатов КТА пациентов с аневризмой брюшной аорты из университетского госпиталя Gachon (Корея), размеченных врачом-рентгенологом с опытом работы более 10 лет.
Сравнение полученного набора данных с аналогами, обнаруженными в открытом доступе, позволяет выделить ряд его преимуществ и недостатков. К достоинствам разработанного набора данных можно отнести:
- объём (100 результатов КТА), превышающий известные аналоги;
- уникальная одновременная представленность случаев нормы, аневризмы, кальциноза и тромбоза брюшной аорты;
- многоцентровой характер включённых исследований;
- верификация первичной разметки с привлечением эксперта;
- ручная попиксельная разметка области интереса;
- открытое декларирование популяционных данных, сценария разметки и требований к разметчикам.
Ограничения исследования
К недостаткам разработанного набора данных и ограничениям исследования следует отнести небольшое количество патологических признаков, включённых в набор данных. Несмотря на наличие ограничений, его можно использовать для решения целого комплекса задач — от классификации различных вариантов атеросклеротического поражения брюшной аорты до антропоморфного моделирования аорты с различными типами поражения её стенки.
ЗАКЛЮЧЕНИЕ
С помощью программного обеспечения открытого доступа Slicer 3D и оригинальной ранее разработанной методике сегментации подготовлен набор данных, содержащий 100 результатов КТА с сегментацией брюшной аортой для случаев нормы, расширения, тромбоза и кальциноза. Набор данных представлен в открытом доступе и его возможно использовать для задач разработки и тестирования алгоритмов искусственного интеллекта, а также для антропоморфного трёхмерного моделирования брюшной аорты.
ДОПОЛНИТЕЛЬНАЯ ИНФОРМАЦИЯ
Источник финансирования. Данная статья подготовлена авторским коллективом в рамках научно-исследовательской и опытноконструкторской работы «Разработка программного обеспечения для автоматического формирования наборов данных КТ-исследований сердечно-сосудистой системы с подавлением контрастирования для обучения и тестирования алгоритмов на основе искусственного интеллекта», (ЕГИСУ: № 123031500002-1) в соответствии с Приказом от 21.12.2022 № 1196 «Об утверждении государственных заданий, финансовое обеспечение которых осуществляется за счёт средств бюджета города Москвы государственным бюджетным (автономным) учреждениям подведомственным Департаменту здравоохранения города Москвы, на 2023 год и плановый период 2024 и 2025 годов» Департамента здравоохранения города Москвы.
Раскрытие интересов. Авторы декларируют отсутствие отношений, деятельности и интересов (личных, профессиональных или финансовых), связанных с третьими лицами (коммерческими, некоммерческими, частными), интересы которых в свою очередь могут быть затронуты содержанием рукописи.
Вклад авторов. М.Р. Коденко — методология исследования, обзор литературы, анализ данных, написание и редактирование статьи; Ю.А. Васильев — планирование исследования и экспертный анализ текста статьи; Д.В. Гатин, А.В. Соловьёв, Е.П. Ясакова — разметка данных и редактирование текста статьи; А.В. Гусева — редактирование текста статьи; Р.В. Решетников — экспертный анализ и редактирование текста статьи. Все авторы одобрили рукопись (версию для публикации), а также согласились нести ответственность за все аспекты работы и гарантировали, что вопросы, связанные с точностью или добросовестностью любой части работы, будут должным образом рассмотрены и решены.
ADDITIONAL INFORMATION
Funding source. This article was prepared by a group of authors as a part of the research and development effort titled "Development of software for automated generation of data sets containing synthetic native-phase CT studies to train and validate AI algorithms", (USIS No. 123031500002-1) in accordance with the Order No. 1196 dated December 21, 2022 "On approval of state assignments funded by means of allocations from the budget of the city of Moscow to the state budgetary (autonomous) institutions subordinate to the Moscow Health Care Department, for 2023 and the planned period of 2024 and 2025" issued by the Moscow Health Care Department.
Disclosure of interests. The authors declare that they have no relationships, activities or interests (personal, professional or financial) with third parties (commercial, non-commercial, private) whose interests may be affected by the content of the article, as well as no other relationships, activities or interests over the past three years that must be reported.
Authors’ contribution. M.R. Kodenko: research methodology, literature review, data analysis, writing and editing the article; Yu.A. Vasilev: research planning and expert analysis of the article text; D.V. Gatin, A.V. Solovev, E.P. Yasakova: data segmentation and editing the article; A.V. Guseva editing the article; R.V. Reshetnikov: expert analysis and editing the article. Thereby, all authors provided approval of the version to be published and agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
1 Aorta segmentation [Internet]. В: Kaggle; 2018–2024. Режим доступа: https://www.kaggle.com/datasets/licethyaneth/aorta-segmentation Дата обращения: 22.05.2024.
2 AAAtester computer vision project [Internet]. В: Roboflow; 2022–2024. Режим доступа: https://universe.roboflow.com/new-workspace-xlgzg/aaatester Дата обращения: 22.05.24.
3 Единый радиологический информационный сервис [Internet]. B: Единая медицинская информационно-аналитическая система города Москвы; 2020–2023. Режим доступа: https://telemedai.ru/proekty/edinyj-radiologicheskij-informacionnyj-servis_2020 Дата обращения: 21.11.2023.
4 3D Slicer image computing platform. В: 3D Slicer [Internet]. 2005–2022. Режим доступа: https://slicer.org/ Дата обращения: 11.09.22.
5 Presize: precision based sample size calculation [Internet]. B: The Swiss Clinical Trial Organisation; 2021–2022. Режим доступа: https://shiny.ctu.unibe.ch/app_direct/presize/ Дата обращения: 07.05.2022.
6 Descriptive Statistics in Excel; [около 14 страниц]. В: Statistics By Jim [Internet]. 2021–2024. Режим доступа: https://statisticsbyjim.com/basics/descriptive-statistics-excel/ Дата обращения: 22.04.2024.
7 Набор данных компьютерно-томографической ангиографии с признаками кальциноза, тромбоза, дилатации и аневризмы, и содержащий сегментацию просвета и стенки брюшного отдела аорты [Internet]. Москва: Научно-практический клинический центр диагностики и телемедицинских технологий Департамента здравоохранения города Москвы; 2024–2024. Режим доступа: https://mosmed.ai/datasets/nabor-dannih-kompyuterno-tomograficheskoi-angiografii-s-priznakami-kaltsinoza-tromboza-dilatatsii-i-anevrizmi-i-soderzhaschii-segmentatsiyu-prosveta-i-stenki-bryushnogo-otdela-aorti/ Дата обращения: 20.04.2024.
8 Dataset Search [Internet]. В: Google; 2018–2024. Режим доступа: https://datasetsearch.research.google.com/ Дата обращения: 22.03.2024.
9 Datasets [Internet]. В: Kaggle; 2021–2024. Режим доступа: https://www.kaggle.com/datasets Дата обращения: 22.03.2024.
10 Explore the Roboflow Universe [Internet]. В: Roboflow; 2021–2023. Режим доступа: https://universe.roboflow.com/ Дата обращения: 22.03.2024.
11 Labeled image datasets for computer vision [Internet]. В: Images.cv; –2024. Режим доступа: https://images.cv/ Дата обращения: 22.03.2024.
12 ITK-SNAP Home [Internet]. В: ITK-SNAP; 2020–2024. Режим доступа: http://www.itksnap.org/pmwiki/pmwiki.php Дата обращения 22.03.2024.
About the authors
Maria R. Kodenko
Research and Practical Clinical Center for Diagnostics and Telemedicine Technologies; Bauman Moscow State Technical University
Author for correspondence.
Email: m.r.kodenko@yandex.ru
ORCID iD: 0000-0002-0166-3768
SPIN-code: 5789-0319
Cand. Sci. (Engineering)
Russian Federation, Moscow; MoscowYuriy A. Vasilev
Research and Practical Clinical Center for Diagnostics and Telemedicine Technologies
Email: VasilevYA1@zdrav.mos.ru
ORCID iD: 0000-0002-5283-5961
SPIN-code: 4458-5608
MD, Cand. Sci. (Medicine)
Russian Federation, MoscowAlexander V. Solovev
Research and Practical Clinical Center for Diagnostics and Telemedicine Technologies; Morozov Children's City Clinical Hospital
Email: SolovevAV10@zdrav.mos.ru
ORCID iD: 0000-0003-4485-2638
SPIN-code: 9654-4005
Russian Federation, Moscow; Moscow
Denis V. Gatin
Research and Practical Clinical Center for Diagnostics and Telemedicine Technologies
Email: GatinDV@zdrav.mos.ru
ORCID iD: 0000-0002-6218-3012
SPIN-code: 2256-3564
Russian Federation, Moscow
Elena P. Yasakova
Research and Practical Clinical Center for Diagnostics and Telemedicine Technologies
Email: YasakovaEP@zdrav.mos.ru
ORCID iD: 0000-0003-0315-5502
SPIN-code: 1047-4692
MD, Cand. Sci. (Medicine)
Russian Federation, MoscowAnastasia V. Guseva
Research and Practical Clinical Center for Diagnostics and Telemedicine Technologies; Bauman Moscow State Technical University
Email: GusevaAV13@zdrav.mos.ru
ORCID iD: 0009-0006-1787-4726
SPIN-code: 2778-3820
Russian Federation, Moscow; Moscow
Roman V. Reshetnikov
Research and Practical Clinical Center for Diagnostics and Telemedicine Technologies
Email: ReshetnikovRV1@zdrav.mos.ru
ORCID iD: 0000-0002-9661-0254
SPIN-code: 8592-0558
Cand. Sci. (Physics and Mathematics)
Russian Federation, MoscowReferences
- Kumar DS, Bhat V, Gadabanahalli K, Kalyanpur A. Spectrum of abdominal aortic disease in a tertiary health care setup: MDCT based observational study. J Clin Diagn Res. 2016;10(11):TC24–TC29. doi: 10.7860/JCDR/2016/21373.8928
- Russian Society of Angiologists and Vascular Surgeons. Abdominal aortic aneurysm: clinical guidelines [Internet]. Moscow: Russian Society of Angiologists and Vascular Surgeons; 2022 [cited 2022 Apr 7]. (In Russ.) Available from: https://angiolsurgery.org/library/recommendations/2022/aneurysm/recommendation.pdf
- Baliyan V, Shaqdan K, Hedgire S, Ghoshhajra B. Vascular computed tomography angiography technique and indications. Cardiovascular Diagnosis and Therapy. 2019;9(S1):S14S27. doi: 10.21037/CDT.2019.07.04 EDN: IPZHHC
- Alowais ShA, Alghamdi SS, Alsuhebany N, et al. Revolutionizing healthcare: the role of artificial intelligence in clinical practice. BMC Medical Education. 2023;23(1):689. doi: 10.1186/s12909-023-04698-z EDN: AJSDXW
- Ueda D, Kakinuma T, Fujita SH, et al. Fairness of artificial intelligence in healthcare: review and recommendations. Japanese Journal of Radiology. 2023;42(1):3–15. doi: 10.1007/s11604-023-01474-3 EDN: WQQDIA
- Shchupakova AN, Litvyakov AM. Characteristics of atherosclerotic lesion of the abdominal aorta and its unpaired visceral branches in patients with chronic abdominal ischemia. Terapevticheskii arkhiv. 2004;79(6):70–74. EDN: OJZUCJ
- Radl L, Jin YU, Pepe A, et al. AVT: Multicenter aortic vessel tree CTA dataset collection with ground truth segmentation masks. Data in Brief. 2022;40:107801. doi: 10.1016/j.dib.2022.107801 EDN: PEOYKJ
- Imran M, Kreds JR, Gopu VRR, et al. CIS-UNet: Multi-class segmentation of the aorta in computed tomography angiography via context-aware shifted window self-attention. Computerized Medical Imaging and Graphics. 2024;118:102470. doi: 10.1016/j.compmedimag.2024.102470
- Fantazzini A, Esposito M, Finotello A, et al. 3D automatic segmentation of aortic computed tomography angiography combining multi-view 2D convolutional neural networks. Cardiovascular Engineering and Technology. 2020;11(5):576–586. doi: 10.1007/s13239-020-00481-z EDN: FHKUXK
- Jung Y, Kim S, Kim J, et al. Abdominal aortic thrombus segmentation in postoperative computed tomography angiography images using Bi-directional cnvolutional long short-term memory architecture. Sensors. 2022;23(1):175. doi: 10.3390/s23010175 EDN: SGCHXK
- Norgeot B, Quer G, Beaulieu-Jones BK, et al. Minimum information about clinical artificial intelligence modeling: the MI-CLAIM checklist. Nature Medicine. 2020;26(9):1320–1324. doi: 10.1038/s41591-020-1041-y EDN: NRQASJ
- Vasilev YuA, Arzamasov KM, Vladzymyrskyy AV, et al. Preparing a dataset for training and testing software based on artificial intelligence technology: a training manual. Moscow: Moscow Center for Diagnostics and Telemedicine; 2023. (In Russ.) EDN: OGKFGM
- Tymkovich MYu, Avruninn OG, Semenets VV. Using DICOM images in medical systems. Technical Electrodynamics. 2012;(thematic issue):178–183. (In Russ.)
- Li X, Morgan P, Ashburner J, et al. The first step for neuroimaging data analysis: DICOM to NIfTI conversion. Journal of Neuroscience Methods. 2016;264:47–56. doi: 10.1016/j.jneumeth.2016.03.001
- Kodenko MR, Vasilev YuA, Vladzymyrskyy AV. Segmentation of arterial vessels based on CT angiography data using 3D Slicer software: Guidelines. Moscow: Moscow Center for Diagnostics and Telemedicine; 2024. (In Russ.) EDN: CYLZQL
- Kodenko MR, Makarova TA. Preparation of abdominal computed tomography data set for patients with abdominal aortic aneurysm. Digital Diagnostics. 2023;4(1S):90–92. doi: 10.17816/DD430355 EDN: SIUWRL
- Riley RD, Snell KIE, Archer L, et al. Evaluation of clinical prediction models (part 3): calculating the sample size required for an external validation study. BMJ. 2024;384:e074821. doi: 10.1136/bmj-2023-074821
- Hazra A. Using the confidence interval confidently. Journal of Thoracic Disease. 2017;9(10):4124–4129. doi: 10.21037/jtd.2017.09.14
- Vasilev YuA, Vladzymyrskyy AV, Arzamasov KM, et al. Computer vision in radiation diagnostics: the first stage of the Moscow experiment. [Internet]. Moscow: Izdatel'skie resheniya; 2022 [cited 2024 Apr 22]. Available from: https://telemedai.ru/biblioteka-dokumentov/kompyuternoe-zrenie-v-luchevoj-diagnostike-pervyj-etap-moskovskogo-eksperimenta
- Certificate of state registration of the database N 2024621990/ 08.05.2024. Byul. N 5. Vasilev YuА, Kodenko МR, Solovev AV, et al. A computed tomography angiography dataset showing calcification, thrombosis, dilation, and aneurysm, and containing segmentation of the lumen and wall of the abdominal aorta. Available from: https://elibrary.ru/download/elibrary_67262583_69739205.PDF (In Russ.) EDN: FOPTBQ
- Guseva AV, Kodenko MR. Anthropomorphic abdominal aortic phantoms for computed tomography angiography. Digital Diagnostics. 2024;5(1S):27–29. doi: 10.17816/DD626820 EDN: BMDJUN
- Lesage D, Angelini ED, Bloch I, Funka-Lea G. A review of 3D vessel lumen segmentation techniques: models, features and extraction schemes. Medical image analysis. 2009;13(6):819–845. doi: 10.1016/j.media.2009.07.011
Supplementary files