Category PI-RADS 3: the role of texture analysis in prostate cancer risk stratification (a systematic review)
- Authors: Tyan A.S.1, Kаrmаzаnovsky G.G.1,2, Karelskaya N.A.1, Kondratyev E.V.1, Gritskevich A.А.1, Kalinin D.V.1, Kovalev A.D.1, Baeva A.I.1
-
Affiliations:
- A.V. Vishnevsky National Medical Research Center of Surgery
- The Russian National Research Medical University named after N.I. Pirogov
- Issue: Vol 6, No 1 (2025)
- Pages: 33-45
- Section: Systematic reviews
- Submitted: 22.06.2024
- Accepted: 18.07.2024
- Published: 25.03.2025
- URL: https://jdigitaldiagnostics.com/DD/article/view/633500
- DOI: https://doi.org/10.17816/DD633500
- ID: 633500
Cite item
Full Text
Abstract
BACKGROUND: Prostate changes classified as PI-RADS 3 are a clinical situation requiring diagnostic accuracy and minimization of invasive procedures. Exploring the potential value of texture analysis in magnetic resonance imaging for prostate cancer risk stratification is critical in modern medical diagnostics.
AIM: To systematize and analyze current data on the application of texture analysis for prostate cancer risk stratification in patients with PI-RADS 3 and evaluate its diagnostic significance in differentiating clinically significant from clinically insignificant prostate cancer.
MATERIALS AND METHODS: Articles published in the last 7 years were selected and analyzed from research reference and analytical databases (Medline and Scopus) using search engines (PubMed, Google Scholar, and eLibrary). Keywords related to texture analysis and radiomics regarding prostate cancer diagnosis and risk stratification were used.
RESULTS: Analysis of the selected publications showed that machine learning and texture analysis significantly enhance the diagnostic accuracy of prostate cancer. These methods allow for more accurate risk stratification and determination of the actual need for biopsy, potentially leading to a reduction in unnecessary invasive procedures.
CONCLUSION: Texture analysis potentially enhances diagnostic accuracy in cases of prostate gland changes classified as PI-RADS 3. However, further research focused on standardizing techniques and conducting multicenter clinical trials is required for its widespread clinical application.
Full Text
ОБОСНОВАНИЕ
Результаты последних исследований, в частности A.C. Westphalen и соавт. [1], продемонстрировали значительную вариабельность прогностичности положительного результата системы PI-RADS v2.1 (Prostate Imaging-Reporting and Data System) в диагностике клинически значимого рака предстательной железы (РПЖ). Именно поэтому возрастает интерес к текстурному анализу изображений магнитно-резонансной томографии (МРТ). Опубликованные данные подчёркивают необходимость разработки и внедрения стандартизированных протоколов и методик, включая применение текстурного анализа для минимизации разночтений и повышения диагностической точности, что особенно важно в случае изменений предстательной железы категории PI-RADS 3, что подразумевает среднюю вероятность наличия клинически значимого РПЖ — сомнительное, присутствие неоднозначное [2–4].
За последние годы текстурный анализ, представляющий собой извлечение большого числа количественных признаков из медицинских изображений, стал перспективным направлением в диагностике и стратификации риска РПЖ [5–7]. С помощью алгоритмов машинного обучения возможен анализ данных для идентификации скрытых паттернов его клинически значимой формы. Они могут способствовать оценке агрессивности опухоли и её реакции на терапию, а также прогнозированию клинических исходов у пациентов [7–11]. Ключевым преимуществом текстурного анализа является возможность получения значимой диагностической информации без необходимости проведения инвазивной процедуры, такой как биопсия, что особенно актуально для случаев с высокой степенью диагностической неопределённости [12, 13].
Широкое распространение текстурный анализ получил в области диагностики клинически значимого РПЖ в рамках системы оценки PI-RADS , особенно категории 3 [3]. Пациенты с РПЖ при наличии очагов категории PI-RADS 3 находятся в серой зоне, где риск выявления его клинически значимой формы недостаточно высок для немедленной биопсии, но и недостаточно низок для отказа от дальнейшего медицинского наблюдения без дополнительных исследований. Это вызывает серьёзные проблемы при ведении данной группы пациентов, поскольку необходимо найти баланс между гипер- или гиподиагностикой и стремлением уменьшить количество инвазивных процедур [2, 14].
В данном контексте текстурный анализ предлагает новые возможности для улучшения точности диагностики клинически значимого РПЖ при наличии очагов категории PI-RADS 3, позволяя более точно стратифицировать риски и выявлять пациентов, нуждающихся в проведении биопсии [13, 15]. Использование сложных алгоритмов для анализа изображений МРТ может выявить нюансы, недоступные визуальной оценке, что позволит принять правильное клиническое решение [2, 13, 16]. В связи с возрастающей актуальностью использования текстурного анализа в диагностике клинически значимого РПЖ мы решили оценить потенциал этого метода в отношении повышения диагностической точности и рассмотреть возможные направления исследований в данной области.
Несмотря на наличие ряда исследований, посвящённых применению текстурного анализа в диагностике РПЖ, включая категорию PI-RADS 3, выявлена значительная вариабельность в результатах и отсутствие стандартизации методов [2–5, 11, 27]. Это подчёркивает необходимость дальнейших исследований для повышения точности диагностики. Настоящее исследование направлено на уточнение роли текстурного анализа в стратификации риска и дифференциации клинически значимого и незначимого РПЖ, что способствует улучшению диагностической точности и снижению неопределённости в интерпретации результатов.
ЦЕЛЬ
Систематизация и анализ современных данных о применении текстурного анализа для стратификации риска РПЖ у пациентов с категорией PI-RADS 3, а также оценка его диагностической значимости в дифференциации клинически значимого и незначимого РПЖ.
МАТЕРИАЛЫ И МЕТОДЫ
В рамках настоящего систематического обзора выполнен всесторонний поиск литературы для оценки роли текстурного анализа в диагностике РПЖ в случае наличия очагов категории PI-RADS 3. Поиск публикаций осуществляли в базах данных реферативной и аналитической информации о научных исследованиях (Medline, Scopus) с использованием поисковых систем PubMed, Google Scholar, и eLibrary. Применяли следующие ключевые слова: «рак предстательной железы» («prostate cancer»), «PI-RADS 3», «текстурный анализ» («texture analysis»), «радиомика» («radiomics»), «магнитно-резонансная томография» («magnetic resonance imaging»), «МРТ» («MRI»), «клинически значимый рак предстательной железы» («clinically significant prostate cancer»).
Критерии включения:
- публикации, описывающие оригинальные исследования в области использования текстурного анализа или радиомики в контексте диагностики РПЖ в случае наличия очагов категории PI-RADS 3;
- публикации, прошедшие внутреннюю или внешнюю валидацию и опубликованные за последние 7 лет.
Критерии исключения:
- публикации без доступного полнотекстового варианта;
- статьи в виде конференционного резюме;
- комментарии, редакционные материалы и публикации, не оценивающие диагностическую точность текстурного анализа.
Процесс отбора публикаций в систематический обзор включал несколько этапов:
- первый — два автора независимо проводили первичный анализ заголовков и аннотаций статей, найденных с помощью ключевых слов;
- второй — авторы независимо оценивали полнотекстовые статьи, соответствующие критериям включения;
- третий — при разногласиях использовали консенсусную стратегию или привлекали третьего независимого эксперта для принятия окончательного решения.
РЕЗУЛЬТАТЫ
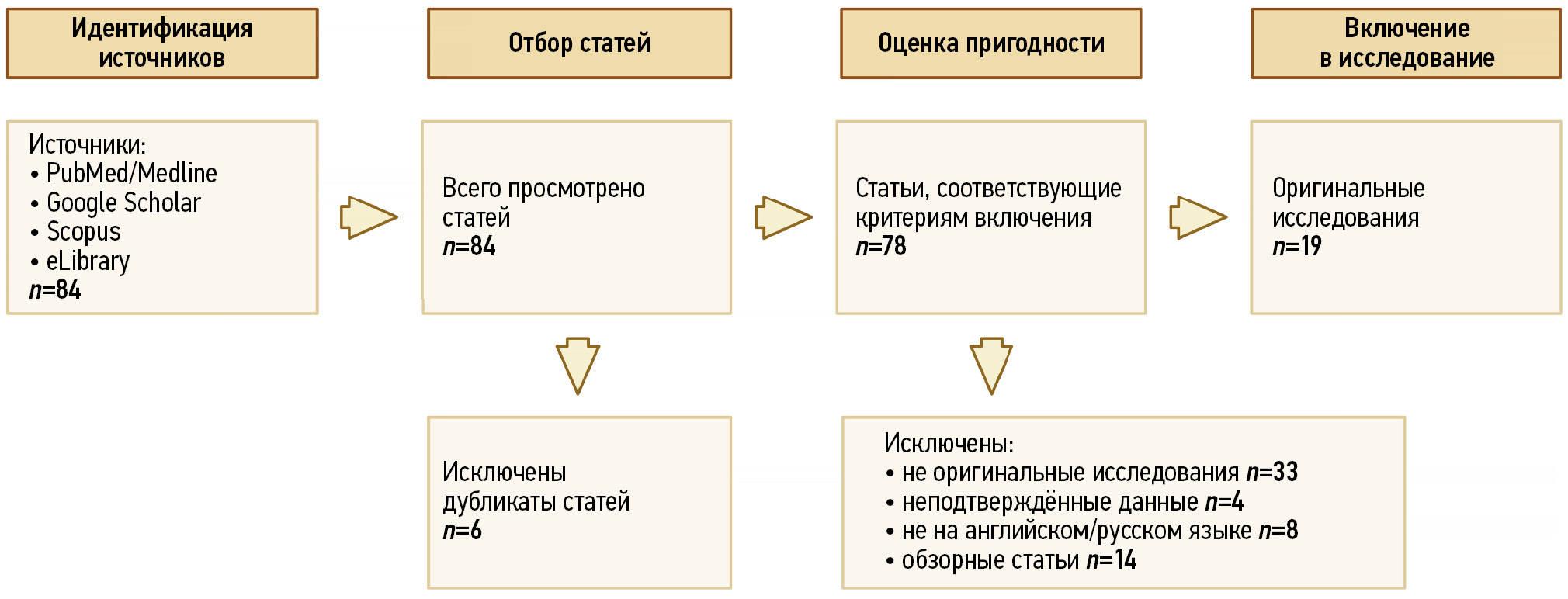
Поиск выявил 85 публикаций, описывающих потенциально релевантные исследования, из которых после детального анализа отобрано 18 статей, соответствующих установленным критериям (рис. 1). Анализ включённых исследований позволил выделить текущие направления и потенциальные возможности повышения эффективности текстурного анализа и радиомики в контексте диагностики РПЖ в случае наличия очагов категории PI-RADS 3 (Приложение 1).
Рис. 1. Схема процесса отбора статей для систематического обзора.
Оценка актуальной проблемы
Комплексный анализ диагностических сложностей, связанных с определением клинически значимого РПЖ в случае наличия очагов категории PI-RADS 3, включает в себя оценку нескольких ключевых аспектов.
Во-первых, в научном сообществе присутствуют существенные разногласия по вопросу дальнейшего ведения пациентов с изменениями предстательной железы категории PI-RADS 3, поскольку имеется риск пропуска малого процента клинически значимых случаев рака. Эта проблема продемонстрирована в работе G. Sonmez и соавт. [17]. Авторы акцентируют внимание на необходимости идентификации предикторов, способствующих выявлению клинически значимого РПЖ у пациентов с концентрацией простатоспецифичного антигена (ПСА) <10 нг/мл и категорией изменений PI-RADS 3.
Во-вторых, существуют методы, помогающие уточнить диагноз, но их считают недостаточно эффективными. Так, I. Hermie и соавт. [13] предлагают комбинацию диагностических методов, а именно: определение содержания ПСА, выполнение мультипараметрической МРТ (мпМРТ) и таргетной биопсии предстательной железы, с целью скрининга рака. Данной стратегии придерживаются и другие авторы, полагая, что она позволит повысить точность диагностики РПЖ в случае наличия очагов категории PI-RADS 3 [15, 18].
В-третьих, A.M. Fang и соавт. [19] в своём исследовании анализируют факторы риска для выявления клинически значимого РПЖ при наличии очагов категории PI-RADS 3. Высокий процент заключений (33%), относящихся к данной категории, свидетельствует о значительных сомнениях рентгенологов в интерпретации результатов, что подчёркивает необходимость поиска дополнительных парадигм поддержки мнения врача [20, 21].
Таким образом, несмотря на наличие определённых подходов к улучшению диагностики РПЖ при наличии изменений категории PI-RADS 3, существует острая необходимость в разработке более точных и надёжных методик. Именно поэтому текстурный анализ может стать перспективным направлением.
Анализ существующих подходов к решению проблемы
Анализ современных научных публикаций в области диагностики клинически значимого РПЖ при наличии очагов категории PI-RADS 3 выявил ряд ключевых аспектов и направлений для дальнейшего изучения.
Одним из предложений является контроль динамики состояния пациентов с РПЖ и изменениями данной категории, однако этот метод несёт в себе риск пропуска его клинически значимой формы. Этот подход демонстрируют в исследованиях, которые описывают важность мониторинга и последующей диагностики, однако он сопряжён с потенциальной опасностью упущения агрессивных форм рака [2, 19, 22].
В соответствии с рекомендациями Международного общества урологической патологии (International Society of Urological Pathology — ISUP) при РПЖ и изменениями категории PI-RADS 3 необходимо проведение дальнейших диагностических мероприятий с целью уточнения диагноза, в частности биопсии [14]. Однако следует отметить, что инвазивное вмешательство сопряжено с риском развития осложнений, требует дополнительного времени на её проведение и возможный период госпитализации, что создаёт дополнительную нагрузку на систему здравоохранения [20]. Таким образом, возникает необходимость в достижении оптимального баланса между точностью диагностики и стремлением к минимизации вмешательства для пациента [2, 23].
Концентрацию ПСА рассматривают в качестве основного биомаркёра для скрининга РПЖ и мониторинга лечения. Тем не менее его оценка не лишена погрешностей, а повышение содержания ПСА не всегда подтверждает наличие клинически значимой формы рака. Так, S. Washino и соавт. [24] в своём исследовании демонстрируют, что увеличение концентрации ПСА не гарантирует наличие РПЖ, однако комплексная оценка данного показателя, результатов мпМРТ и таргетной биопсии позволяют повысить диагностическую точность.
Текстурный анализ представляет собой новаторский метод, способный улучшить точность диагностики, уменьшить риск пропуска клинически значимого РПЖ и облегчить нагрузку на систему здравоохранения благодаря выполнению только обоснованных биопсий [25, 26].
Таким образом, перед современной медициной стоит задача интеграции этих технологий в клиническую практику с целью повышения диагностической точности в отношении подтверждения клинически значимого РПЖ в случае наличия очагов категории PI-RADS 3, сокращая при этом риски для пациентов и снижая потребление ресурсов здравоохранения.
Обзор методов текстурного анализа
Инициативы по интеграции текстурного анализа в медицинскую практику могут быть классифицированы на два основных направления, дифференцируемых в соответствии с первостепенными задачами аналитического процесса.
В первую очередь идентификация и сегментация подозрительных на наличие неоплазии участков. Это направление включает в себя разработку и валидацию алгоритмических решений для автоматизированного выделения участков ткани, подозрительных в отношении опухолевого процесса, с целью повышения диагностической эффективности [27].
Во-вторых, оценка степени агрессивности карциномы предстательной железы. Второе направление сфокусировано на анализе извлечённых данных из томографических изображений с целью предсказания агрессивности опухолевого процесса [6, 7]. Текстурный анализ может извлекать из изображений скрытые паттерны, которые человеческий глаз не способен обнаружить [25, 26]. Они могут коррелировать с генетическими и молекулярными характеристиками опухоли, что позволяет оценить её агрессивность и потенциальное течение [28]. Использование такого подхода может помочь в выборе стратегии лечения, определяя пациентов, нуждающихся в немедленном вмешательстве либо динамическом контроле. Это особенно важно в случае РПЖ при наличии изменений категории PI-RADS 3, где решение о лечении должно балансировать между риском необоснованно агрессивной диагностики и необходимостью своевременной терапии для предотвращения прогрессирования заболевания [3, 25, 29].
В данных направлениях ключевым является разработка и валидация алгоритмов, которые возможно интегрировать в клиническую практику. Это требует тесного сотрудничества между врачами-рентгенологами, врачамионкологами и специалистами в области информационных технологий для обеспечения точности, воспроизводимости и клинической релевантности радиомических моделей [27].
Основные направления инициатив интеграции текстурного анализа можно классифицировать на шесть отдельных подкатегорий (рис. 2).
Рис. 2. Схема направлений исследований по применению текстурного анализа в диагностике рака предстательной железы при наличии очагов категории PI-RADS 3. РПЖ — рак предстательной железы; PI-RADS 3 (Prostate Imaging-Reporting and Data System 3) — категория 3, средняя вероятность (клинически значимый рак сомнителен, присутствие неоднозначное).
Разработка и валидация моделей машинного обучения
Исследования в области разработки и валидации моделей машинного обучения, направленные на выявление клинически значимого РПЖ при наличии очагов категории PI-RADS 3, осуществлены различными исследовательскими группами. В частности, D. Kwon и соавт. [10] разработали классификацию подозрительных поражений с помощью машинного обучения по данным мпМРТ предстательной железы. Данная модель включала элементы трансферного обучения, что способствовало повышению предсказательной точности. Например, использовали свёрточные нейронные сети для извлечения признаков и экстремальные машины обучения с весовыми коэффициентами для классификации. Основная цель данного исследования — разработка алгоритма компьютерной диагностики, основанного на принципах глубокого машинного обучения, для верификации РПЖ.
S.G. Hectors и соавт. [29] разработали и апробировали модель машинного обучения на основе радиомических признаков, извлечённых из Т2-взвешенных изображений, полученных с помощью МРТ, для прогнозирования клинически значимого РПЖ в случае наличия изменений категории PI-RADS 3. Она продемонстрировала высокие показатели чувствительности и специфичности, превосходя верификацию врачами-рентгенологами разного уровня квалификации [29].
Кроме того, P. Jin. и соавт. [16] провели ретроспективное мультицентровое исследование с целью разработки радиомической модели с помощью машинного обучения, предназначенной для прогнозирования доброкачественных и злокачественных новообразований с изменениями предстательной железы категории PI-RADS 3. Это исследование подчеркнуло важность внешней валидации и потенциал для обобщения результатов более широкой популяции.
Ключевыми достижениями исследований являются разработка моделей с высокой точностью, которые превосходно выявляют клинически значимый РПЖ. Это может привести к сокращению количества необоснованных биопсий, что является значительным вкладом в клиническую практику [16, 23, 29].
Однако несмотря на данные достижения, выявлены также определённые недостатки. В частности, модели глубокого машинного обучения обладают ограниченной интерпретируемостью, что затрудняет понимание механизмов их работы и может осложнить широкое внедрение таких технологий в медицинскую практику без дополнительных исследований и адаптации. Кроме того, их эффективность зависит от объёма и качества доступных данных — в случае нехватки или низкого качества наблюдают существенное снижение точности и надёжности моделей, что подчёркивает важность создания крупных и анонимизированных наборов данных для обучения и тестирования таких систем [16, 29, 30].
Анализ текстурных признаков и методологии извлечения данных
G. Penzias и соавт. [31] осуществили анализ морфологической основы текстурных признаков для стратификации РПЖ в соответствии со шкалой Глисона по данным МРТ. Выявлено, что наиболее предсказательной ценностью обладают спектральные текстурные признаки, полученные с помощью фильтров Габора [площадь под кривой (AUC) составляет 0,69] и паттерны формы просвета желёз (AUC составляет 0,75), что подчёркивает значимость изменений в форме и морфологии желёз на тканевом уровне для дифференциации степени РПЖ по данным МРТ. Результаты исследования позволили переосмыслить подход к использованию текстурного анализа для прогнозирования и идентификации клинически значимых форм РПЖ, что может способствовать оптимизации диагностики и стратегий лечения. Основным преимуществом данного исследования является возможность точного неинвазивного определения степени агрессивности опухоли посредством анализа результатов МРТ. Однако существуют сложности в интерпретации радиомических данных, а также зависимость результатов от качества и параметров оборудования, используемого для проведения МРТ.
D. Giambelluca и соавт. [25] оценивали диагностическую эффективность текстурного анализа изображений МРТ предстательной железы в отношении РПЖ в случае изменений категории PI-RADS 3. Авторы продемонстрировали хорошую диагностическую ценность текстурного анализа для его идентификации — получены прогностические модели для выявления РПЖ (по шкале Глисона ≥6 баллов) и его клинически значимой формы (по шкале Глисона ≥7 баллов) со значением AUC от 0,775 до 0,821 и 0,749 до 0,817 соответственно. Разработана прогностическая модель, сочетающая показатель плотности ПСА1 с двумя текстурными признаками, извлечёнными из изображений МРТ, что позволило достичь высокой чувствительности и специфичности при верификации клинически значимого РПЖ — 80 и 76% соответственно. Полученные результаты позволяют повысить эффективность ранней диагностики, что в дальнейшем сокращает количество необоснованных биопсий.
Оценка производительности моделей и их клиническая значимость
В рамках недавнего исследования предложено использование результатов мпМРТ и текстурного анализа для скрининга РПЖ у мужчин с концентрацией ПСА от 4 до 10 нг/мл. Исследование направлено на определение возможности прогнозирования клинически значимого РПЖ на основе текстурных признаков, полученных из изображений мпМРТ, что может избавить пациентов с низким риском заболевания от необходимости проведения биопсии. Эффективность модели зависит от соблюдения стандартов проведения исследований в соответствии с концепцией PI-RADS v2.1, а также требует высокой квалификации специалистов для адекватной интерпретации результатов текстурного анализа [32].
L. Gong и соавт. [33] исследовали применение различных последовательностей бипараметрической МРТ (бпМРТ) предстательной железы для неинвазивной диагностики рака высокой степени злокачественности. Все радиомические модели показали значимые прогностические характеристики, что позволяет улучшить существующие методики скрининга, предоставляя врачам инструменты для более точного определения тактики ведения пациентов с данной патологией. Y. Wang и соавт. [34] также продемонстрировали неинвазивный метод прогнозирования клинически значимого РПЖ. Однако для создания модели с помощью машинного обучения они использовали радиомические особенности изображений МРТ, клинические данные и уровень метилирования промотора PRKY, что позволило достичь высокой прогностической эффективности. Разработка радиомических моделей прогнозирования позволит улучшить точность диагностики клинически значимого РПЖ, что поспособствует более адекватному выбору тактики лечения. Тем не менее данные модели требуют сложной валидации, что приводит к ограничениям в применении из-за различий в технической оснащённости и доступности оборудования для выполнения МРТ [34, 35].
Описанные исследования демонстрируют значительный потенциал использования продвинутых технологий визуализации и анализа данных для улучшения точности диагностики и персонализации подходов к лечению РПЖ. Однако необходимы дальнейшие исследования для подтверждения их клинической эффективности и безопасности.
Оценка воспроизводимости и проведение внешней валидации
Х. Min и соавт. [8] создали радиомическую модель на основе признаков, извлечённых из изображений, полученных с помощью мпМРТ, для дифференциальной диагностики клинически значимых и незначимых форм РПЖ. Следует отметить, что особое внимание уделяли кросс-валидации методов машинного обучения для оценки устойчивости и воспроизводимости модели на неоднородных данных. Применение регуляризованной логистической регрессии и методик кросс-валидации позволило провести объективную оценку эффективности предложенной модели, что является перспективным подходом к улучшению точности диагностики РПЖ. Включение механизмов кросс-валидации способствует повышению надёжности и обобщённости модели, что критически важно для её стабильности и воспроизводимости в различных клинических условиях. Однако несмотря на высокую статистическую надёжность, существует риск переобучения и потенциальные трудности в применении модели без специализированных знаний в области машинного обучения и текстурного анализа.
Интеграция текстурных признаков с клиническими показателями в диагностических моделях
В рамках исследований, проведённых P. Woźnicki и соавт. [9], а также A. Corsi и соавт. [35], осуществлена интеграция текстурных признаков с клиническими параметрами для улучшения диагностики РПЖ в случае наличия изменений категории PI-RADS 3.
Так, P. Woźnicki и соавт. [9] продемонстрировали, что модели машинного обучения, основанные на количественных радиомических признаках, извлечённых из изображений мпМРТ, в сочетании с клиническими характеристиками обладают диагностической ценностью в отношении клинически значимого РПЖ. Этот метод подчёркивает важность радиомической модели для увеличения диагностической эффективности системы PI-RADS v2.1, улучшая качество стратификации риска РПЖ.
A. Corsi и соавт. [35] отмечают важность текстурного анализа в отношении детализации оценок в соответствии с системой PI-RADS v 2.1, особенно при сомнительных результатах МРТ. Предложенная клинико-диагностическая модель, включающая текстурные признаки, значительно повышает диагностическую эффективность, обеспечивая более точное определение стратегий ведения пациентов и уменьшая количество необоснованных биопсий.
Ключевые преимущества этих методов заключены в улучшении точности и специфичности диагностических процедур, что критически важно для оптимизации подходов к лечению. Однако стандартизация и воспроизводимость, а также необходимость в большом объёме данных для обучения и валидации моделей остаются значительными препятствиями для широкого внедрения этих технологий в клиническую практику.
Стратификация риска
Предприняты значительные шаги в направлении улучшения диагностики РПЖ и его стратификации риска с использованием текстурного анализа, направленного на извлечение данных из изображений, полученных с помощью мпМРТ [36–38].
V. Brancato и соавт. [36] оценивали роль радиомических признаков, извлечённых из Т2-взвешенных и постконтрастных изображений, полученных с помощью мпМРТ, для идентификации РПЖ в случае наличия очагов категории PI-RADS 3 и 4. Для извлечения радиомических признаков из медицинских изображений авторы использовали пакет PyRadiomics, с помощью которого проанализировали более 290 признаков, отобранных через тест Вилкоксона и алгоритм mRMR. Разработанные логистические регрессионные модели демонстрировали высокую эффективность в отношении идентификации РПЖ, превосходя систему PI-RADS v2.1 с показателями AUC до 0,89 для категории PI-RADS 4. Исследование подчёркивает важность радиомического анализа с целью улучшения диагностики РПЖ и стратификации его риска.
Y. Hou и соавт. [37] с помощью машинного обучения разработали радиомическую модель для выявления клинически значимого РПЖ при наличии изменений категории PI-RADS 3. Авторы продемонстрировали, что одношаговая радиомическая модель (RML-i) показала высокое значение AUC — 0,89, что свидетельствует о повышении точности стратификации риска. Данная модель представляет собой алгоритм машинного обучения, обрабатывающий весь набор текстурных признаков за один процесс без необходимости предварительного отбора или ручной корректировки признаков. Она автоматически анализирует изображения и выделяет нужные признаки, оптимизируя процесс обучения и увеличивая общую точность предсказаний. Такая модель особенно полезна в клинических условиях, где необходимо быстрое и точное принятие решений на основе сложных медицинских данных.
Y. Zhang и соавт. [38] разработали новую радиомическую номограмму, включающую признаки, извлечённые из изображений мпМРТ, для дифференциальной диагностики клинически значимого и незначимого РПЖ. Номограмма содержит клинические характеристики и радиомические признаки для точной верификации РПЖ у пациентов с концентрацией ПСА 4–10 нг/мл.
Представленные исследования подчёркивают значимость текстурного анализа, направленного на извлечение данных из изображений, полученных с помощью мпМРТ, в улучшении стратификации риска и точности диагностики РПЖ.
Ограничения и перспективы
Несмотря на обнадёживающие результаты исследований, посвящённых изучению текстурного анализа в диагностике РПЖ, существуют работы, подвергающие сомнению его эффективность [39, 40].
Так, W. Krauss и соавт. [40] провели исследование с участием 350 пациентов, которым проводили мпМРТ. Авторы анализировали разнообразные текстурные признаки, извлечённые из изображений мпМРТ, такие как характеристики текстуры и гистограмм, с целью выявления их корреляции с наличием и агрессивностью РПЖ на основе результатов последующих биопсий и клинических исходов. Исследование показало, что несмотря на возможность с помощью текстурного анализа проводить дифференциальную диагностику доброкачественных и злокачественных новообразований, интеграция текстурного анализа с традиционными параметрами МРТ и клиническими данными не привела к значительному повышению прогностической точности. Например, значения AUC для моделей с использованием текстурного анализа не имели отличия от таковых для моделей без его применения.
Подтверждая данные этого исследования, E. Gresser и соавт. [39] также получили аналогичные результаты, демонстрирующие воспроизводимость выводов о потенциале текстурного анализа при использовании данных МРТ. В исследовании участвовало 142 пациента, которым выполнили МРТ предстательной железы в разных медицинских учреждениях. Полученные результаты показали, что значение AUC моделей варьировало от 0,78 до 0,83, подчёркивая высокую диагностическую ценность метода. Однако значительная вариабельность результатов, особенно при использовании различных стратегий коррекции смещения, требует дальнейших исследований с целью улучшения стабильности радиомических измерений. Следует отметить, что несмотря на использование современных методов машинного обучения, интеграция текстурного анализа с традиционными методами диагностики, такими как анализ измеряемого коэффициента диффузии и оценка по системе PI-RADS, не привела к значительному улучшению прогностической точности.
Полученные результаты указывают на то, что текстурные признаки могут иметь различия в зависимости от используемых методов визуализации, что ограничивает их широкое применение в медицинской практике. Кроме того, отсутствие единых алгоритмов текстурного анализа и различия в протоколах получения изображений в разных клиниках затрудняют внедрение радиомических моделей прогнозирования в обычные клинические процессы [39, 40].
Для преодоления существующих ограничений продолжают разрабатывать научные методики, способствующие оптимизации процедур в данной области. Существуют публикации, описывающие стандартизацию процессов и минимизацию ошибок на всех этапах интеграции текстурного анализа в клиническую практику. Так, E.P. Huang и соавт. [41] предлагают 16 критериев для эффективного внедрения текстурного анализа в клинические тесты. Они охватывают весь процесс — от стандартизации методик извлечения признаков до оценки клинической пользы и валидности радиомических тестов. Авторы предлагают несколько ключевых направлений для дальнейшей работы в области радиомики.
- Гармонизация и стандартизация процессов. Основное внимание необходимо уделять гармонизации и стандартизации процессов извлечения текстурных признаков и их анализа. Это включает установление стандартных операционных процедур для получения изображений и извлечения признаков, а также статистического анализа. Авторы подчёркивают важность стандартизация, поскольку она способствует повышению надёжности и воспроизводимости радиомических исследований.
- Разработка и валидация моделей. Рекомендовано разрабатывать радиомические модели с мерами предотвращения переобучения и их последующей валидацией. Это включает применение данных внешней валидации, которые не использованы при разработке модели, или адекватных методов внутренней валидации, если внешние данные недоступны.
- Итеративное уточнение и адаптация. Необходимо подчеркнуть важность мониторинга изменений в радиомических тестах, которые могут происходить из-за обновлений аппаратного и программного обеспечения. Какие-либо изменения, не связанные с дрейфом данных, такие как применение теста в другой популяции пациентов или по другим показаниям, должны требовать возвращения к предыдущим этапам разработки и валидации моделей.
Эти направления представляют собой комплексный подход к улучшению и расширению областей применения текстурного анализа в клинической практике с целью увеличения числа полезных радиомических исследований [5, 41].
ОБСУЖДЕНИЕ
При рассмотрении аспектов применения текстурного анализа в диагностике РПЖ в случае наличия очагов категории PI-RADS 3 необходимо обозначить проблемы, которые, вероятно, невозможно решить с помощью этого метода в обозримом будущем.
Стандартизация методов получения изображений и анализа. В различных исследованиях подчёркивают зависимость текстурного анализа от качества и согласованности медицинских изображений. Следует отметить, что в разных медицинских учреждениях используют различные протоколы сканирования и параметры МРТ, что влияет на воспроизводимость результатов анализа. Стандартизация методов получения изображений по-прежнему остаётся крупной проблемой, которая требует дальнейших усилий для разработки универсальных руководств и протоколов [1].
Интерпретируемость моделей машинного обучения. Несмотря на то что модели глубокого обучения на основе текстурного анализа показывают многообещающие результаты в классификации и предсказании РПЖ, их интерпретируемость остаётся сложной задачей. Это означает, что понимание нюансов принятия определённых диагностических решений может быть неочевидным. Данное ограничение затрудняет внедрение в клиническую практику таких технологий, поскольку медицинским специалистам важно понимать систему работы диагностических инструментов [4].
Переобучение и обобщённость моделей. Существует риск переобучения моделей машинного обучения, особенно в случае их обучения на ограниченных или не полностью репрезентативных данных. Переобучение ведёт к тому, что модели хорошо работают только на данных, используемых для обучения. Обобщённость таких моделей на более широкую популяцию пациентов остаётся под вопросом, требуя дополнительных усилий для внешней валидации и тестирования на разнообразных наборах данных [6].
Нерешённые вопросы оценки степени агрессивности и стадирования рака. Текстурный анализ предоставляет возможность выявлять наличие опухоли, однако точное определение стадии и агрессивности опухолевого процесса остаётся сложной задачей. Некоторые биологические аспекты опухоли не отражены в изменениях текстуры изображений, что ограничивает способность метода точно стратифицировать риски и определять наиболее подходящее лечение [5].
Эти проблемы подчёркивают сложность и многогранность задачи диагностики РПЖ с помощью текстурного анализа и машинного обучения, указывая на необходимость дальнейших научных исследований и разработок в этой области.
ЗАКЛЮЧЕНИЕ
Текстурный анализ, особенно в сочетании с методами машинного обучения, представляет собой многообещающий метод в рамках диагностики РПЖ при наличии очагов категории PI-RADS 3. Применение этих технологий способствует улучшению точности диагностических моделей, позволяя более точно стратифицировать риски и определять необходимость биопсии, тем самым потенциально сокращая количество ненужных инвазивных процедур.
Однако существуют и значительные проблемы, связанные с внедрением этих технологий в клиническую практику. Основные трудности связаны c высокой вариабельностью текстурных признаков, зависящих от технических характеристик оборудования, используемого для проведения МРТ, и параметров сканирования, что может уменьшать возможность применения радиомических моделей в различных условиях и снижать их воспроизводимость. Важно отметить необходимость стандартизации методик извлечения и анализа данных для обеспечения согласованности результатов на различных платформах и в разных клинических условиях.
С целью преодоления этих препятствий необходимо сосредоточить будущие исследования в области разработки стандартизированных протоколов для текстурного анализа, учитывающих разнообразие технических параметров аппаратуры и последовательностей для проведения МРТ. Кроме того, важной задачей является создание крупных аннотированных баз данных, что позволит улучшить обучение и валидацию моделей машинного обучения. Эти шаги помогут повысить клиническую значимость текстурного анализа в диагностике РПЖ.
Таким образом, несмотря на значительный потенциал текстурного анализа для улучшения диагностики РПЖ, наше понимание того, как наилучшим образом использовать эти технологии в клинической практике, ещё находится на начальном этапе развития.
ДОПОЛНИТЕЛЬНАЯ ИНФОРМАЦИЯ
Приложение 1. Резюме исследований по применению текстурного анализа в диагностике клинически значимого рака предстательной железы. doi: 10.17816/DD633500-4272791
Источник финансирования. Авторы заявляют об отсутствии внешнего финансирования при проведении исследования.
Раскрытие интересов. Авторы заявляют об отсутствии отношений, деятельности и интересов (личных, профессиональных или финансовых), связанных с третьими лицами (коммерческими, некоммерческими, частными), интересы которых могут быть затронуты содержанием статьи, а также иных отношений, деятельности и интересов за последние три года, о которых необходимо сообщить.
Вклад авторов. А.С. Тян — анализ данных, написание текста статьи; Е.В. Кондратьев — концепция и дизайн исследования; Г.Г. Кармазановский, Н.А. Карельская, А.А. Грицкевич — редактирование и утверждение итогового варианта текста рукописи, консультативная поддержка; Д.В. Калинин — консультативная поддержка; А.А. Ковалёв, А.И. Баева — анализ данных. Все авторы одобрили рукопись (версию для публикации), а также согласились нести ответственность за все аспекты работы и гарантировали, что вопросы, связанные с точностью или добросовестностью любой части работы, будут должным образом рассмотрены и решены.
ADDITIONAL INFORMATION
Appendix 1. Summary of studies on the application of texture analysis in the diagnosis of clinically significant prostate cancer. doi: 10.17816/DD633500-4272791
Funding source. This study was not supported by any external sources of funding.
Disclosure of interests. The authors declare that they have no relationships, activities or interests (personal, professional or financial) with third parties (commercial, non-commercial, private) whose interests may be affected by the content of the article, as well as no other relationships, activities or interests over the past three years that must be reported.
Authors’ contribution. A.S. Tyan: data analysis, manuscript writing; E.V. Kondratyev: research concept and design; G.G. Karmazanovsky, N.A. Karelskaya, A.A. Gritskevich: manuscript editing and final approval, advisory support; D.V. Kalinin: advisory support; A.A. Kovalev, A.I. Baeva: data analysis. Thereby, all authors provided approval of the version to be published and agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
1 Плотность ПСА — показатель, используемый в диагностике заболеваний предстательной железы, включая рак. Представляет собой отношение концентрации ПСА в сыворотке крови к объёму предстательной железы.
About the authors
Alexandra S. Tyan
A.V. Vishnevsky National Medical Research Center of Surgery
Author for correspondence.
Email: tyan_a_s@staff.sechenov.ru
ORCID iD: 0009-0007-4193-7413
SPIN-code: 9110-9827
Russian Federation, Moscow
Grigory G. Kаrmаzаnovsky
A.V. Vishnevsky National Medical Research Center of Surgery; The Russian National Research Medical University named after N.I. Pirogov
Email: karmazanovsky@ixv.ru
ORCID iD: 0000-0002-9357-0998
SPIN-code: 5964-2369
MD, Dr. Sci. (Medicine), Professor, academician of the Russian Academy of Sciences
Russian Federation, Moscow; MoscowNatalia A. Karelskaya
A.V. Vishnevsky National Medical Research Center of Surgery
Email: karelskaya.n@yandex.ru
ORCID iD: 0000-0001-8723-8916
SPIN-code: 9921-1430
MD, Cand. Sci. (Medicine)
Russian Federation, MoscowEvgeny V. Kondratyev
A.V. Vishnevsky National Medical Research Center of Surgery
Email: kondratev@ixv.ru
ORCID iD: 0000-0001-7070-3391
SPIN-code: 2702-6526
MD, Cand. Sci. (Medicine)
Russian Federation, MoscowAlexander А. Gritskevich
A.V. Vishnevsky National Medical Research Center of Surgery
Email: grekaa@mail.ru
ORCID iD: 0000-0002-5160-925X
SPIN-code: 2128-7536
MD, Dr. Sci. (Medicine), Professor
Russian Federation, MoscowDmitry V. Kalinin
A.V. Vishnevsky National Medical Research Center of Surgery
Email: dmitry.v.kalinin@gmail.com
ORCID iD: 0000-0001-6247-9481
SPIN-code: 5563-5376
MD, Cand. Sci. (Medicine)
Russian Federation, MoscowAlexander D. Kovalev
A.V. Vishnevsky National Medical Research Center of Surgery
Email: aledmikov@yandex.ru
ORCID iD: 0009-0001-9944-0473
Russian Federation, Moscow
Anastasiya I. Baeva
A.V. Vishnevsky National Medical Research Center of Surgery
Email: nastya.baeva.2016@mail.ru
ORCID iD: 0000-0003-3747-7411
Russian Federation, Moscow
References
- Westphalen AC, McCulloch CE, Anaokar JM, et al. Variability of the Positive Predictive Value of PI-RADS for Prostate MRI across 26 Centers: Experience of the Society of Abdominal Radiology Prostate Cancer Disease-focused Panel. Radiology. 2020;296(1):76–84. doi: 10.1148/radiol.2020190646 EDN: FJNEKA
- Wadera A, Alabousi M, Pozdnyakov A, et al. Impact of PI-RADS Category 3 lesions on the diagnostic accuracy of MRI for detecting prostate cancer and the prevalence of prostate cancer within each PI-RADS category: a systematic review and meta-analysis. The British Journal of Radiology. 2020;94(1118):20191050. doi: 10.1259/bjr.20191050
- Satei AM, Kaur M, MacLean J, Hakim B. Review of clinically significant cancer in lesions labeled PI-RADS 3 on MRI using PI-RADS Version 2.1. Applied Radiology. 2023;(1 Suppl.):13–19.
- Barkovich EJ, Shankar PR, Westphalen AC. A systematic review of the existing prostate imaging reporting and data system version 2 (PI-RADSv2) literature and subset meta-analysis of PI-RADSv2 categories stratified by Gleason Scores. American Journal of Roentgenology. 2019;212(4):847–854. doi: 10.2214/AJR.18.20571
- Stanzione A, Gambardella M, Cuocolo R, et al. Prostate MRI radiomics: a systematic review and radiomic quality score assessment. European Journal of Radiology. 2020;129:109095. doi: 10.1016/j.ejrad.2020.109095 EDN: GUEKIR
- Bao J, Qiao X, Song Y, et al. Prediction of clinically significant prostate cancer using radiomics models in real-world clinical practice: a retrospective multicenter study. Insights into Imaging. 2024;15(1):68. doi: 10.1186/s13244-024-01631-w EDN: RFQGMA
- Gelezhe PB, Blokhin IA, Semenov SS, Caruso D. Magnetic resonance imaging radiomics in prostate cancer radiology: what is currently known? Digital Diagnostics. 2021;2(4):441–452. doi: 10.17816/DD70170 EDN: FFFGWI
- Min X, Li M, Dong D, et al. Multi-parametric MRI-based radiomics signature for discriminating between clinically significant and insignificant prostate cancer: cross-validation of a machine learning method. European Journal of Radiology. 2019;115:16–21. doi: 10.1016/j.ejrad.2019.03.010
- Woźnicki P, Westhoff N, Huber T, et al. Multiparametric MRI for prostate cancer characterization: combined use of radiomics model with PI-RADS and clinical parameters. Cancers (Basel). 2020;12(7):1767. doi: 10.3390/cancers12071767 EDN: DMCTFH
- Kwon D, Reis IM, Breto AL, et al. Classification of suspicious lesions on prostate multiparametric MRI using machine learning. Journal of Medical Imaging. 2018;5(3):034502. doi: 10.1117/1.JMI.5.3.034502
- He M, Cao Y, Chi C, et al. Research progress on deep learning in magnetic resonance imaging–based diagnosis and treatment of prostate cancer: a review on the current status and perspectives. Frontiers in Oncology. 2023;13:1189370. doi: 10.3389/fonc.2023.1189370 EDN: FEZIDE
- Chen Z, Li Z, Dou R, et al. Personalized optimization of systematic prostate biopsy core number based on mpMRI radiomics features: a large–sample retrospective analysis. BMC Cancer. 2025;25(1):116. doi: 10.1186/s12885-024-13391-3
- Hermie I, Van Besien J, De Visschere P, et al. Which clinical and radiological characteristics can predict clinically significant prostate cancer in PI-RADS 3 lesions? A retrospective study in a high-volume academic center. European Journal of Radiology. 2019;114:92–98. doi: 10.1016/j.ejrad.2019.02.031
- Epstein JI, Egevad L, Amin MB, et al. The 2014 International Society of Urological Pathology (ISUP) consensus conference on gleason grading of prostatic carcinoma. American Journal of Surgical Pathology. 2016;40(2):244–252. doi: 10.1097/PAS.0000000000000530
- Gulin GA, Zyryanov AV, Rubtsova NA, et al. Multiparametric magnetic resonance imaging and combined prostate biopsy: opportunities, advantages and pitfalls. Medical Visualization. 2021;25(2):138–152. doi: 10.24835/1607-0763-1023 EDN: MSXQIH
- Jin P, Shen J, Yang L, et al. Machine learning-based radiomics model to predict benign and malignant PI-RADS v2.1 category 3 lesions: a retrospective multi-center study. BMC Medical Imaging. 2023;23(1):47. doi: 10.1186/s12880-023-01002-9 EDN: QFWKOM
- Sonmez G, Tombul ST, Demirtas T, Demirtas A. Clinical factors for predicting malignancy in patients with PSA <10 ng/mL and PI-RADS 3 lesions. Asia-Pacific Journal of Clinical Oncology. 2020;17(2):e94–e99. doi: 10.1111/ajco.13347 EDN: ZVXTXF
- Vasilev AV, Mishchenko AV, Kadyrleev RA, et al. Cognitive mpMRI/TRUS biopsy of the prostate with using strain elastography. Medical Visualization. 2019;23(2):100–108. doi: 10.24835/1607-0763-2019-2-100-108 EDN: UPQEEH
- Fang AM, Shumaker LA, Martin KD, et al. Multi-institutional analysis of clinical and imaging risk factors for detecting clinically significant prostate cancer in men with PI-RADS 3 lesions. Cancer. 2022;128(18):3287–3296. doi: 10.1002/cncr.34355 EDN: HHUNTK
- Park KJ, Choi SH, Lee JS, et al. Risk stratification of prostate cancer according to PI-RADS® version 2 categories: meta-analysis for prospective studies. The Journal of Urology. 2020;204(6):1141–1149. doi: 10.1097/ju.0000000000001306 EDN: ASHPIN
- Önder Ö, Ayva M, Yaraşır Y, et al. Long-term follow-up results of multiparametric prostate MRI and the prognostic value of PI-RADS: a single-center retrospective cohort study. Diagnostic and Interventional Radiology. 2024;30(3):139–151. doi: 10.4274/dir.2023.232414
- Gromov AI, Kapustin VV. Usage of PI-RADS v2.1 system for prostate MRI: a practical approach. Medical Visualization. 2019;23(3):107–125. doi: 10.24835/1607-0763-2019-3-107-125 EDN: AHBXBT
- Gupta RT, Mehta KA, Turkbey B, Verma S. PI-RADS: Past, present, and future. Journal of Magnetic Resonance Imaging. 2020;52(1):33–53. doi: 10.1002/jmri.26896
- Washino S, Okochi T, Saito K, et al. Combination of prostate imaging reporting and data system (PI-RADS) score and prostate-specific antigen (PSA) density predicts biopsy outcome in prostate biopsy naïve patients. BJU International. 2017;119(2):225–233. doi: 10.1111/bju.13465
- Giambelluca D, Cannella R, Vernuccio F, et al. PI-RADS 3 lesions: role of prostate MRI texture analysis in the identification of prostate cancer. Current Problems in Diagnostic Radiology. 2021;50(2):175–185. doi: 10.1067/j.cpradiol.2019.10.009 EDN: KZLCMA
- Kаrmаzаnovsky GG, Shantarevich MY, Stashkiv VI, Revishvili AS. Reproducibility of CT and MRI texture features of hepatocellular carcinoma. Medical Visualization. 2023;27(3):84–93. doi: 10.24835/1607-0763-1372 EDN: BVBFNX
- Ferro M, de Cobelli O, Musi G, et al. Radiomics in prostate cancer: an up-to-date review. Therapeutic Advances in Urology. 2022;14:17562872221109020. doi: 10.1177/17562872221109020
- Fischer S, Tahoun M, Klaan B, et al. A Radiogenomic approach for decoding molecular mechanisms underlying tumor progression in prostate cancer. Cancers. 2019;11(9):1293. doi: 10.3390/cancers11091293
- Hectors SJ, Chen C, Chen J, et al. Magnetic resonance imaging radiomics-based machine learning prediction of clinically significant prostate cancer in equivocal PI-RADS 3 lesions. Journal of Magnetic Resonance Imaging. 2021;54(5):1466–1473. doi: 10.1002/jmri.27692 EDN: SGBOWM
- Magoulianitis V, Yang J, Yang Y, et al. PCa-RadHop: a transparent and lightweight feed-forward method for clinically significant prostate cancer segmentation. Computerized Medical Imaging and Graphics. 2024;116:102408. doi: 10.1016/j.compmedimag.2024.102408 EDN: XEOIXK
- Penzias G, Singanamalli A, Elliott R, et al. Identifying the morphologic basis for radiomic features in distinguishing different Gleason grades of prostate cancer on MRI: preliminary findings. PLOS ONE. 2018;13(8):e0200730. doi: 10.1371/journal.pone.0200730
- Qi Y, Zhang S, Wei J, et al. Multiparametric MRI-Based radiomics for prostate cancer screening with PSA in 4–10 ng/mL to reduce unnecessary biopsies. Journal of Magnetic Resonance Imaging. 2019;51(6):1890–1899. doi: 10.1002/jmri.27008 EDN: EINUYX
- Gong L, Xu M, Fang M, et al. Noninvasive prediction of high-grade prostate cancer via biparametric MRI radiomics. Journal of Magnetic Resonance Imaging. 2020;52(4):1102–1109. doi: 10.1002/jmri.27132 EDN: EMVNVC
- Wang Y, Liu W, Chen Z, et al. A noninvasive method for predicting clinically significant prostate cancer using magnetic resonance imaging combined with PRKY promoter methylation level: a machine learning study. BMC Medical Imaging. 2024;24(1):60. doi: 10.1186/s12880-024-01236-1 EDN: RASMDD
- Corsi A, De Bernardi E, Bonaffini PA, et al. Radiomics in PI-RADS 3 multiparametric MRI for prostate cancer identification: literature models re-Implementation and proposal of a clinical–radiological model. Journal of Clinical Medicine. 2022;11(21):6304. doi: 10.3390/jcm11216304 EDN: GGECVG
- Brancato V, Aiello M, Basso L, et al. Evaluation of a multiparametric MRI radiomic-based approach for stratification of equivocal PI-RADS 3 and upgraded PI-RADS 4 prostatic lesions. Scientific Reports. 2021;11(1):643. doi: 10.1038/s41598-020-80749-5 EDN: HAJXQY
- Hou Y, Bao ML, Wu CJ, et al. A radiomics machine learning-based redefining score robustly identifies clinically significant prostate cancer in equivocal PI-RADS score 3 lesions. Abdominal Radiology. 2020;45(12):4223–4234. doi: 10.1007/s00261-020-02678-1 EDN: JHWPET
- Zhang Y, Chen W, Yue X, et al. Development of a novel, multi-parametric, MRI-based radiomic nomogram for differentiating between clinically significant and insignificant prostate cancer. Frontiers in Oncology. 2020;10(FEB):888. doi: 10.3389/fonc.2020.00888 EDN: QTDXSC
- Gresser E, Schachtner B, Stüber AT, et al. Performance variability of radiomics machine learning models for the detection of clinically significant prostate cancer in heterogeneous MRI datasets. Quantitative Imaging in Medicine and Surgery. 2022;12(11):4990–5003. doi: 10.21037/qims-22-265 EDN: JIFGJJ
- Krauss W, Frey J, Heydorn Lagerlöf J, et al. Radiomics from multisite MRI and clinical data to predict clinically significant prostate cancer. Acta Radiologica. 2023;65(3):307–317. doi: 10.1177/02841851231216555 EDN: VVMHZC
- Huang EP, O’Connor JPB, McShane LM, et al. Criteria for the translation of radiomics into clinically useful tests. Nature Reviews Clinical Oncology. 2022;20(2):69–82. doi: 10.1038/s41571-022-00707-0 EDN: XNGIOB
Supplementary files